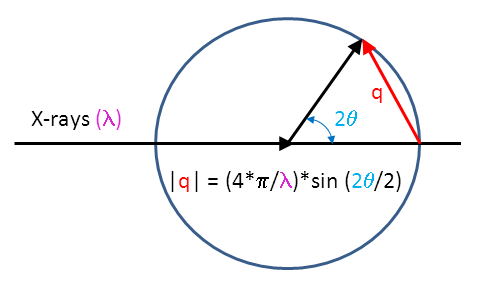
Plot of the form factor P(q) of a sphere with R = 30 Å. An example of a... | Download Scientific Diagram
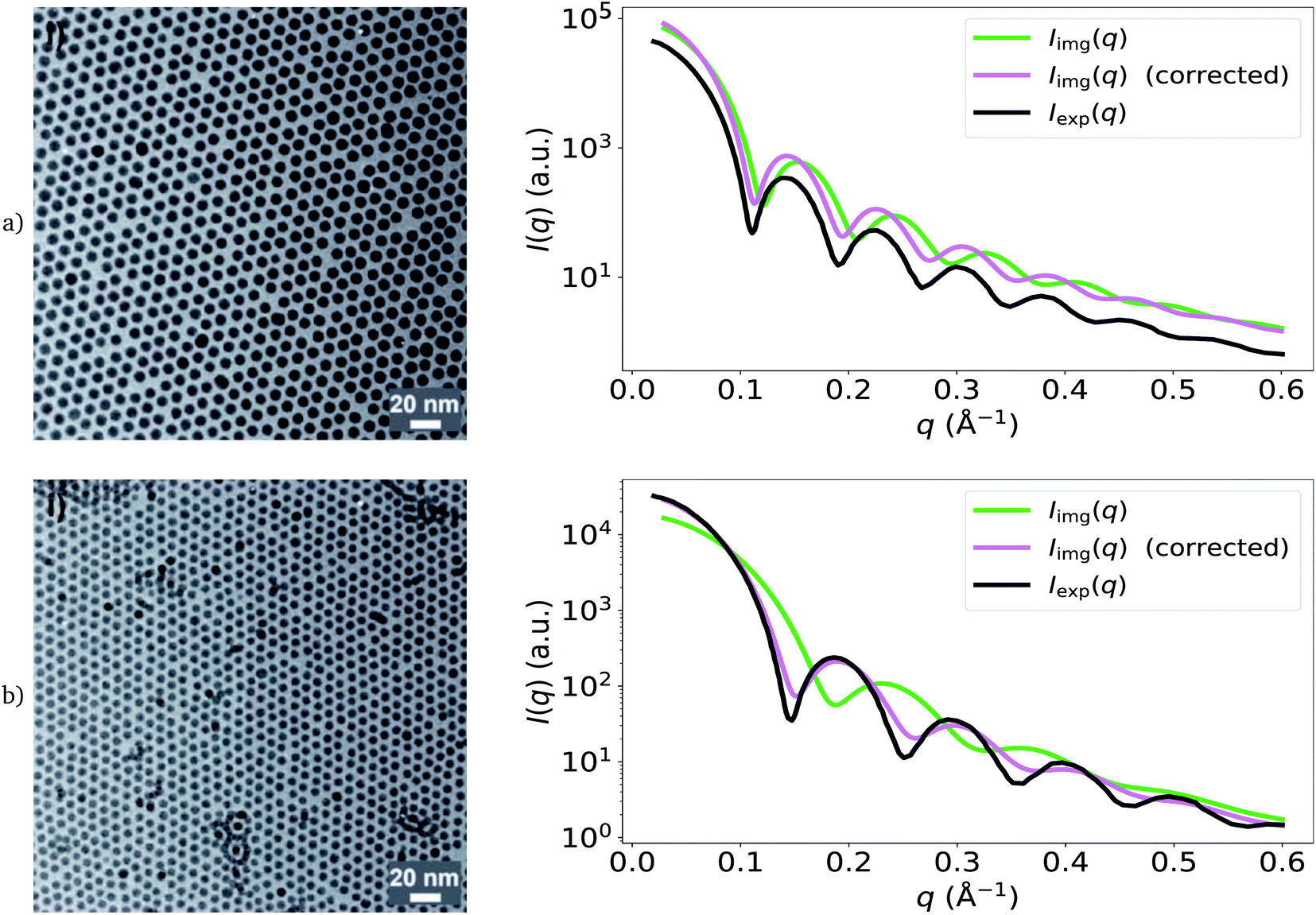
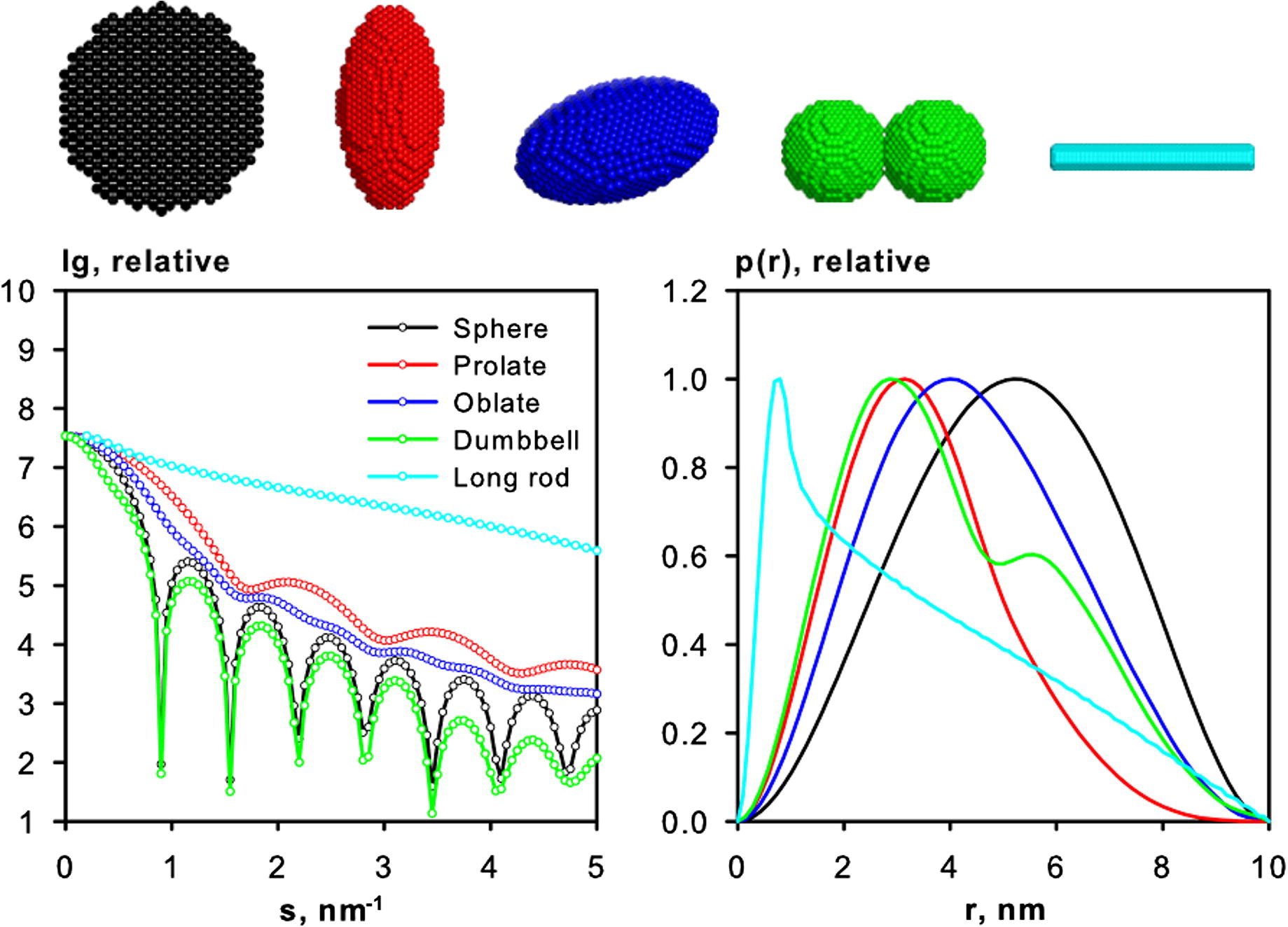
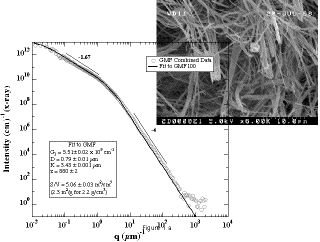
Calculating small-angle scattering intensity functions from electron-microscopy images - RSC Advances (RSC Publishing) DOI:10.1039/D2RA00685E
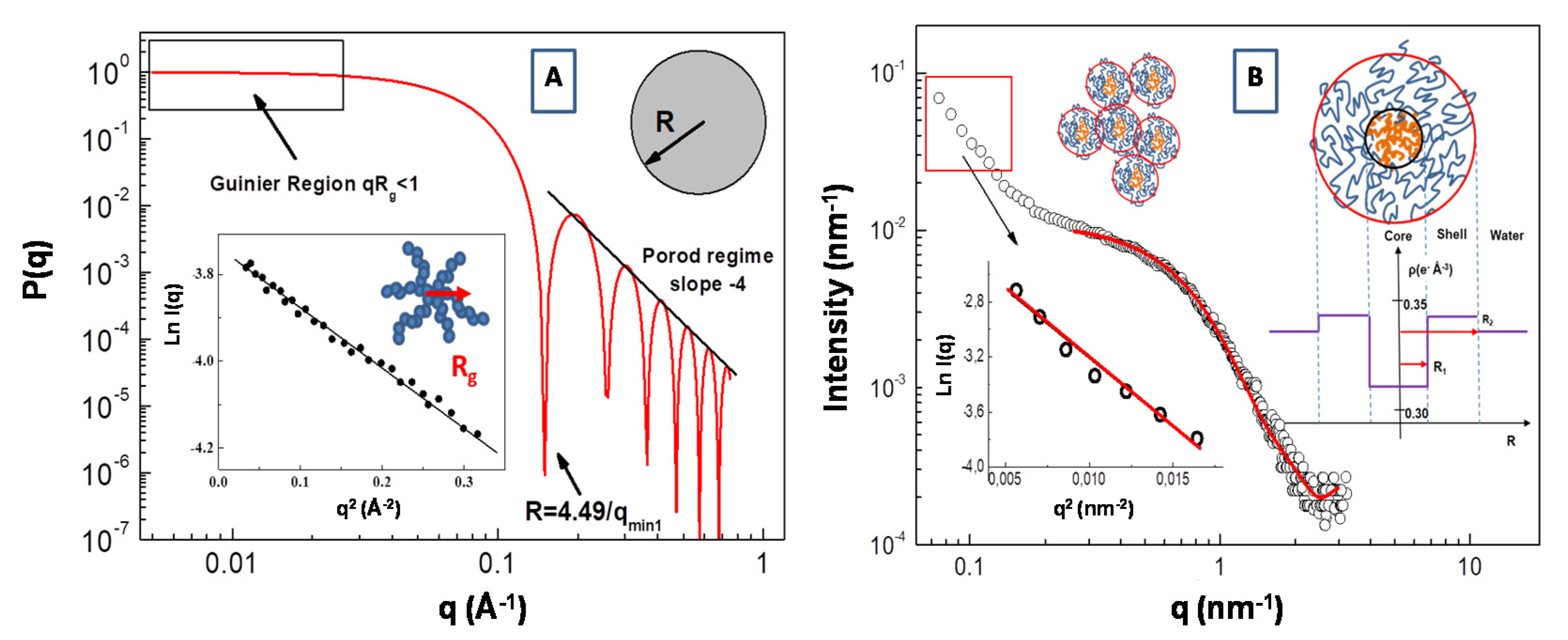
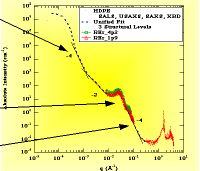
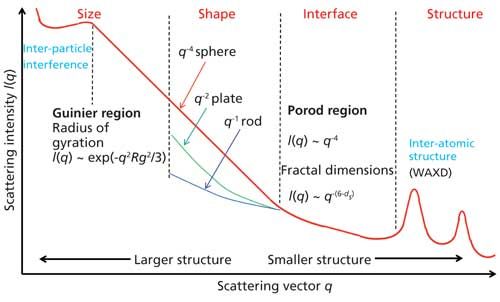
Molecules | Free Full-Text | Structural Characterization of Biomaterials by Means of Small Angle X-rays and Neutron Scattering (SAXS and SANS), and Light Scattering Experiments

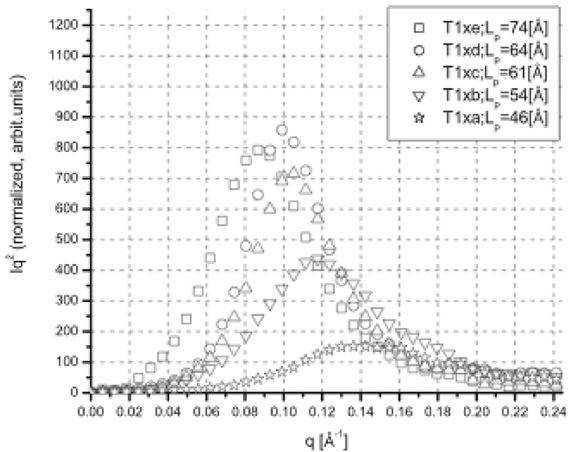
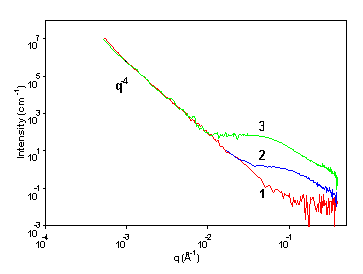
Small‐Angle X‐Ray Scattering Studies of Block Copolymer Nano‐Objects: Formation of Ordered Phases in Concentrated Solution During Polymerization‐Induced Self‐Assembly - Rymaruk - 2021 - Angewandte Chemie International Edition - Wiley Online Library

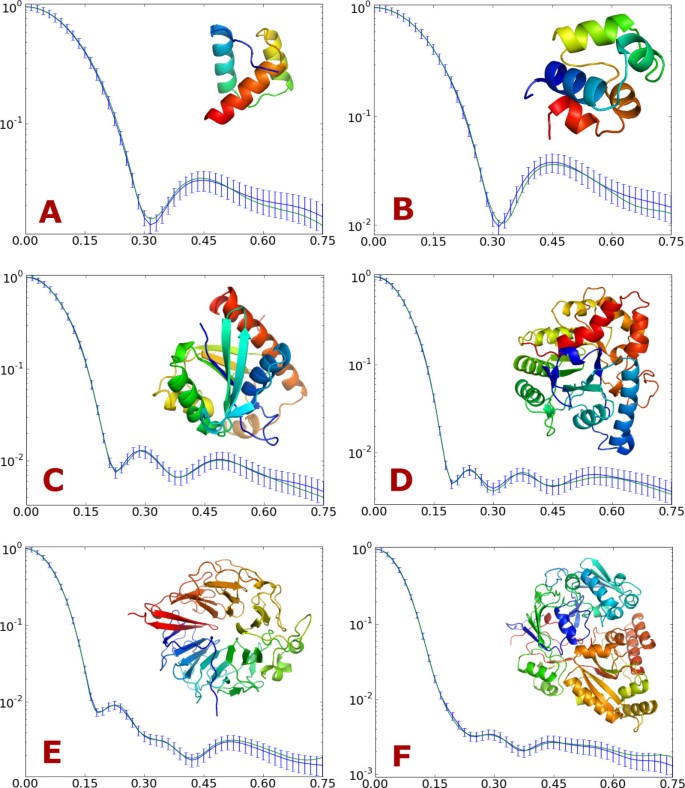
SAXS-guided Enhanced Unbiased Sampling for Structure Determination of Proteins and Complexes | Scientific Reports
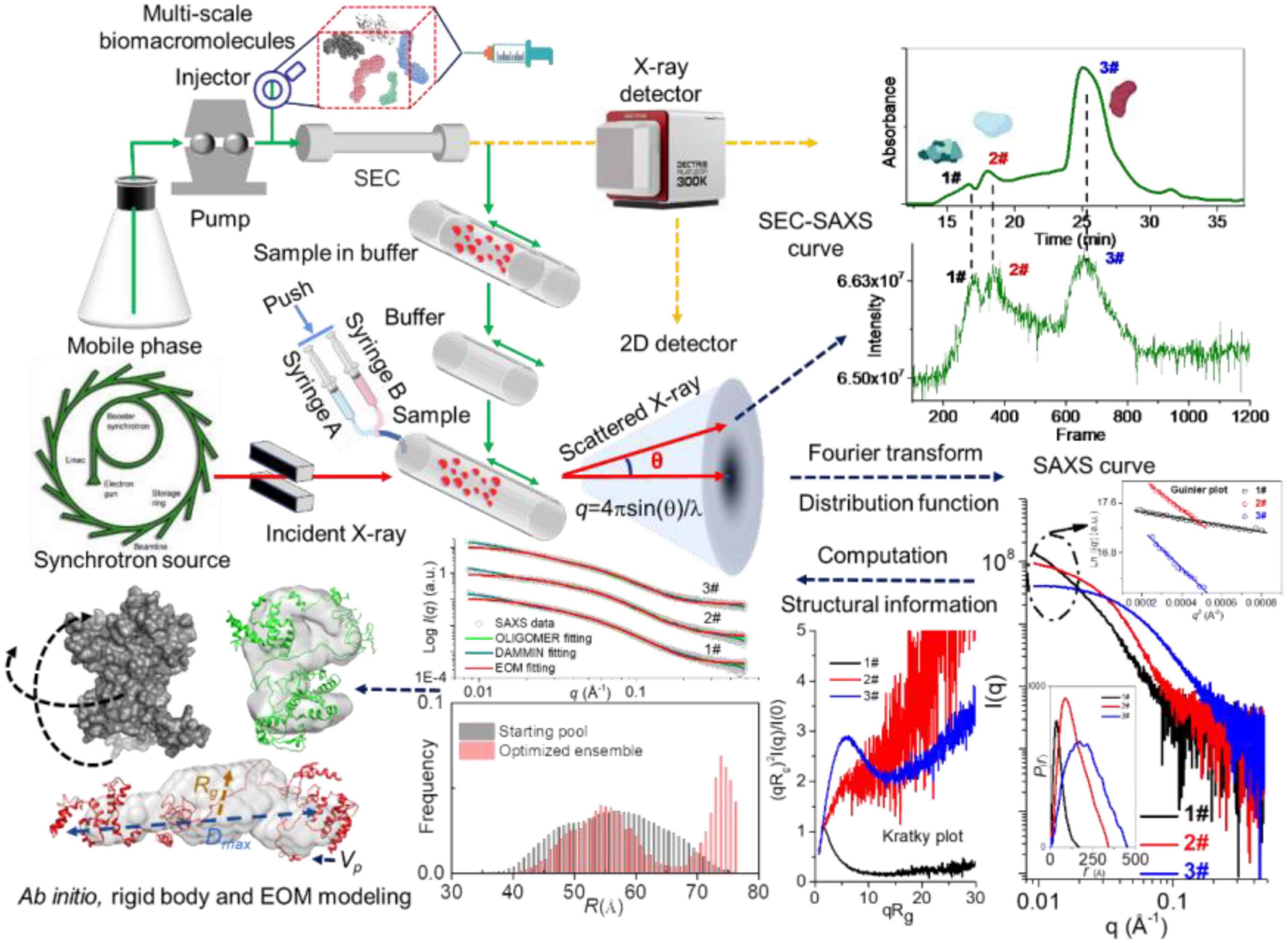
Frontiers | Recent advances in structural characterization of biomacromolecules in foods via small-angle X-ray scattering

Simulation-Based Fitting of Protein-Protein Interaction Potentials to SAXS Experiments: Biophysical Journal